The International Union for Conservation of Nature, publisher of the infamous Red List of endangered species, estimates that 25% of all assessed species are threatened with extinction. Currently genetic data is not included in IUCN species assessment, but what if species at risk could be identified using genomic data? One group of Purdue Forestry and Natural Resources affiliated researchers compiled whole genome sequences, habitat availability data and life-history trait data (such as body mass and diet) for 68 bird species to find out how biological and environmental conditions have shaped genomic diversity and why genomic diversity differs among species in order to provide insights for future conservation efforts.
The results of this research were published in the Proceedings of the Royal Society B Biological Sciences article titled “Life History Traits and Habitat Availability Shape Genomic Diversity in Birds: Implications for Conservation". The paper was co-authored by Drs. Anna Brüniche-Olsen, Ken Kellner (MS 2012, PhD 2015), professor of genetics Andrew DeWoody and professor Jerrold Belant of the Global Wildlife Conservation Center at the State University of New York College of Environmental Science and Forestry (SUNY-ESF), who is now a professor in the department of Fisheries and Wildlife at Michigan State University.
Brüniche-Olsen, a former postdoctoral research assistant in the DeWoody Lab, is currently the Carlsberg postdoctoral fellow at the University of Copenhagen. Kellner, who was named as the early career honoree of the 2021 FNR Chase S. Osborn Award in Wildlife Conservation, is a research scientist and biometrician at the Global Wildlife Conservation Center.
“The goal was to investigate if we could use whole genome data and environmental niche modelling to identify species at risk of extinction,” Brüniche-Olsen explained. “We found that genomic diversity is positively correlated with the amount of habitat available to the species. The more habitat available to a species, the bigger the population size and amount of genomic diversity. Furthermore, we found that life-history traits such as body mass and diet influence genomic diversity, with larger bodied species having lower genomic diversity and species at the lower trophic levels having higher genomic diversity.
“Species categorized as threatened by IUCN have significantly lower genomic diversity. This is of great concern as genomic diversity is fundamental for whether a species will be able to adapt to environmental change. Our findings illustrate that whole genome sequence data can be use to identify species at risk of extinction and may be informative in conservation management.”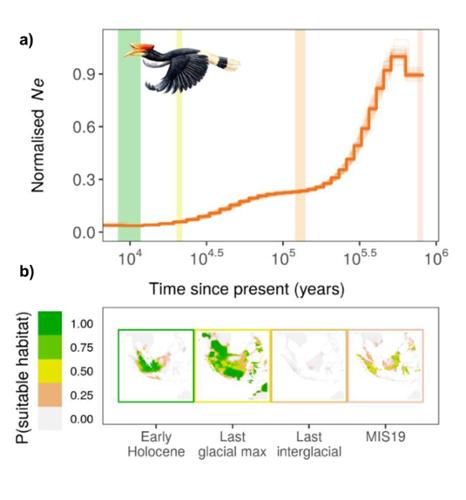
After collecting the whole genome sequence data on 68 bird species, the researchers estimated overall genomic diversity by inferring population sizes for each species over the past million years and reconstructing the amount of available habitat for each species at three periods during that same time span.
Kellner developed ecological niche models (ENMs) for each bird species. ENMs use reported locations of a species along with present-day climate data (temperature, precipitation, etc.) to identify the range of climate conditions that represent suitable habitat for the species.
After building a model for each species, researchers applied the models to climate data from several periods in the past to generate predications about how much suitable habitat a species may have had during those periods. They then looked for relationships between the genomics data and the amount of predicted suitable habitat in the past.
Their conclusions showed that threatened bird species have lower genomic diversity and that genomic diversity is in part driven by habitat availability over time. They also found that the effects of diet on genomic diversity were the largest among life-history traits. Finally, results suggested that when assessing species genomic variation, using species of similar body mass and diet as reference provide context for assessing levels of genomic diversity.
This publication marks the continuation of the trio’s research in the DeWoody lab, where they previously addressed how increased use of whole genome sequencing could find overall patterns that could be used to inform conservation.
“I’ve been really fortunate to work with this outstanding team of scientists,” DeWoody said.
Past publications include “Runs of homozygosity have utility in mammalian conservation and evolutionary studies", which investigated how inbreeding level in mammals scales with life-history traits, and “Island area, body size and demographic history shape genomic diversity in Darwin’s finches and related tanagers", which looked at how extent of habitat shaped genomic diversity in this iconic species complex.
“Our research highlights the importance of conserving habitat for species persistence,” Brüniche-Olsen said. “Identifying species at risk at an early stage means that conservation management can be implemented, funds can be directed to management of the focal species and the chance of mitigating the risk factors will be better.
“All of this research helps broaden the field of conservation genetics and adds to the discussion of how we can use DNA sequencing data in conservation of species.”





